|
 |
"Bald Eagle" <cre### [at] netscape net> wrote:
> Hi Thomas,
>
> Buckminsterfullerene (BF) is quite interesting - I remember clearly when they
> were discovered in 1985 - it was a big deal, even though I was only in HS and
> didn't understand why (yet).
>
> From a didactic point, you should have started with basic bonding,
> Hund/Aufbau/orbital-filling, VSEPR, and orbital hybridization to show how atoms
> bond, why they adopt the shapes that they do, and explain why certain atoms have
> the number of bonds that they do. And of course, since we're talking about
> Buckminsterfullerene, you should probably mention aromaticity, and what makes BF
> interesting and why its discovery was so important, and why 3 people won the
> Nobel prize for its discovery...
> (Even if you only do so VERY briefly. You could also just talk about it and
> provide links in your video description / GitHub)
> I know, I know....
>
> Although I'm personally happy that you covered the topic, I think you might
> scare off some people by diving right into the geometry of dihedral angles, etc.
>
> "Reflecting all the time I invested in this molecule, it seems pretty easy to
> model, ..." Maybe. I haven't tried your way yet, and when I looked at your
> code on GitHub, it's 1134 lines. I think it might be time to introduce arrays
> and macros, and the blessings of shorter code. :)
> Also, if you're thinking of modeling larger molecules, maybe #read from ASCII
> CSV file would be a good topic to cover in advance.
> (I found an .xyz file at:
>
https://www.researchgate.net/profile/Marco-De-La-Pierre/publication/308325208_Coordinates_of_nn_fullerenes_with_n_1-1
0_
> xyz_format/data/57e0cdc608ae3f2d793ebb43/fulle-xyz.zip
> )
>
> Since Buckminsterfullerene is a truncated icosahedron, I used the description of
> the vertices here:
> https://en.wikipedia.org/wiki/Truncated_icosahedron
> and once I puzzled out the meaning of the "even permutations" part, I think I
> have all the vertices plotted. (without having them all connected it's hard to
> tell)
> (I was really hoping that someone had invented some sort of amazing parametric
> equation that would give the vertex locations as the result! :D )
>
> Maybe once I have it all worked out and connected with bonds, I can compare the
> positions of atoms in each model.
>
> It's interesting that you do almost all of your work with blobs.
> First, blobs were developed by James Blinn to model DNA for Carl Sagan's
> _Cosmos_. So, there's some wicked cool history there.
> One of my very first renders in the early 2000's was a DNA double-helix! :)
>
> Second, jr was just asking about blobs, and I found a lot of interesting
> information about how metaballs can be implemented, with different shapes other
> than spheres, and using different smoothing functions. Pixar's Renderman has
> some pretty impressive stuff in that area.
>
> I noticed that you had some interesting creases in the contours of your blobs -
> I'm wondering if we can improve on POV-Ray's stock blob implementation...
>
> Overall, a nice video, showing how to build up a complex structure, layer by
> layer, and using symmetry to simplify the task.
>
> Another interesting way to model the 2nd half of the structure would be to use
> the symmetry through the center of inversion. And maybe the lower half could be
> simplified into 5 sections that could be copied by rotation.
> http://www.creative-science.org.uk/c60group.html
>
> I'm looking forward to seeing what your future ideas for the series are!
>
> - Bill
Hi Bill,
thanks for all the constructive comments - also in the subsequent discussion! I
will not cover everything here, but rather implement a few things in later
videos/scripts.
Referring to basic chemistry, I am a bit unsure to which extent people are
really interested in this. Anyway, the fullerene was not thought as the starting
structure of my chemistry part and I will use the initial structures to talk a
bit more about theory. Maybe above all we should discuss what we are actually
visualizing when presenting atoms and bonds...
Admittedly my script is quite lengthy and I could have shortened by using loops
and arrays. I already introduced loops and arrays in other scripts and I will
use them later at other opportunities, but I thought for people unexperienced in
Pov-Ray this lengthy form might be easier to understand. (In addition I wanted
to avoid the work necessary to implement this here...)
I had a look at the wiki file for truncated isoahedrons and, o.k., this is also
possible. Even superior, when it is about elegance/shortness of code. The
advantage of my code is a) that I completely understand the approach (...) and
b) that after this lengthy definition of points I can now address each point
separately, knowing which point is where. So I could take this structure as a
basis for something else. (O.k., currently I have no idea what I actually could
do, in particular since fullerenes are not very reactive I guess...)
Anyway, there are always several pathways to a given problem.
I have to admit that I can't really follow the technical discussion about blobs.
It only came to my mind that blobs are somewhat similar to electron orbitals.
Orbitals as far as I understand are clouds of probability to encounter an
electron with, if I remember correctly, increasing probability towards the
center of the cloud. This is actually quite similar to metaballs. I'm not
claiming, of course that the blobs I have been using are somehow resembling the
actual fullerene orbitals.
It was interesting to look at the read-function - I never used it... I extracted
data for small molecules always by hand from xyz-files. In the case of large
molecules I wrote my own Perl-scripts for transforming pdb-data into pov-files.
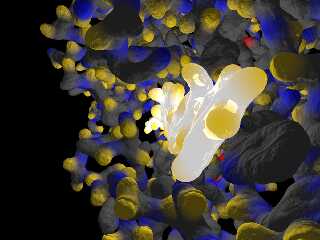
As a teaser for these larger molecules, here comes a detail from the active
center of an endoglucanase with its substrate.
Btw, I apologize for posting so rarely and that it will also take some more time
until my next video...
Thomas net> wrote:
> Hi Thomas,
>
> Buckminsterfullerene (BF) is quite interesting - I remember clearly when they
> were discovered in 1985 - it was a big deal, even though I was only in HS and
> didn't understand why (yet).
>
> From a didactic point, you should have started with basic bonding,
> Hund/Aufbau/orbital-filling, VSEPR, and orbital hybridization to show how atoms
> bond, why they adopt the shapes that they do, and explain why certain atoms have
> the number of bonds that they do. And of course, since we're talking about
> Buckminsterfullerene, you should probably mention aromaticity, and what makes BF
> interesting and why its discovery was so important, and why 3 people won the
> Nobel prize for its discovery...
> (Even if you only do so VERY briefly. You could also just talk about it and
> provide links in your video description / GitHub)
> I know, I know....
>
> Although I'm personally happy that you covered the topic, I think you might
> scare off some people by diving right into the geometry of dihedral angles, etc.
>
> "Reflecting all the time I invested in this molecule, it seems pretty easy to
> model, ..." Maybe. I haven't tried your way yet, and when I looked at your
> code on GitHub, it's 1134 lines. I think it might be time to introduce arrays
> and macros, and the blessings of shorter code. :)
> Also, if you're thinking of modeling larger molecules, maybe #read from ASCII
> CSV file would be a good topic to cover in advance.
> (I found an .xyz file at:
>
https://www.researchgate.net/profile/Marco-De-La-Pierre/publication/308325208_Coordinates_of_nn_fullerenes_with_n_1-1
0_
> xyz_format/data/57e0cdc608ae3f2d793ebb43/fulle-xyz.zip
> )
>
> Since Buckminsterfullerene is a truncated icosahedron, I used the description of
> the vertices here:
> https://en.wikipedia.org/wiki/Truncated_icosahedron
> and once I puzzled out the meaning of the "even permutations" part, I think I
> have all the vertices plotted. (without having them all connected it's hard to
> tell)
> (I was really hoping that someone had invented some sort of amazing parametric
> equation that would give the vertex locations as the result! :D )
>
> Maybe once I have it all worked out and connected with bonds, I can compare the
> positions of atoms in each model.
>
> It's interesting that you do almost all of your work with blobs.
> First, blobs were developed by James Blinn to model DNA for Carl Sagan's
> _Cosmos_. So, there's some wicked cool history there.
> One of my very first renders in the early 2000's was a DNA double-helix! :)
>
> Second, jr was just asking about blobs, and I found a lot of interesting
> information about how metaballs can be implemented, with different shapes other
> than spheres, and using different smoothing functions. Pixar's Renderman has
> some pretty impressive stuff in that area.
>
> I noticed that you had some interesting creases in the contours of your blobs -
> I'm wondering if we can improve on POV-Ray's stock blob implementation...
>
> Overall, a nice video, showing how to build up a complex structure, layer by
> layer, and using symmetry to simplify the task.
>
> Another interesting way to model the 2nd half of the structure would be to use
> the symmetry through the center of inversion. And maybe the lower half could be
> simplified into 5 sections that could be copied by rotation.
> http://www.creative-science.org.uk/c60group.html
>
> I'm looking forward to seeing what your future ideas for the series are!
>
> - Bill
Hi Bill,
thanks for all the constructive comments - also in the subsequent discussion! I
will not cover everything here, but rather implement a few things in later
videos/scripts.
Referring to basic chemistry, I am a bit unsure to which extent people are
really interested in this. Anyway, the fullerene was not thought as the starting
structure of my chemistry part and I will use the initial structures to talk a
bit more about theory. Maybe above all we should discuss what we are actually
visualizing when presenting atoms and bonds...
Admittedly my script is quite lengthy and I could have shortened by using loops
and arrays. I already introduced loops and arrays in other scripts and I will
use them later at other opportunities, but I thought for people unexperienced in
Pov-Ray this lengthy form might be easier to understand. (In addition I wanted
to avoid the work necessary to implement this here...)
I had a look at the wiki file for truncated isoahedrons and, o.k., this is also
possible. Even superior, when it is about elegance/shortness of code. The
advantage of my code is a) that I completely understand the approach (...) and
b) that after this lengthy definition of points I can now address each point
separately, knowing which point is where. So I could take this structure as a
basis for something else. (O.k., currently I have no idea what I actually could
do, in particular since fullerenes are not very reactive I guess...)
Anyway, there are always several pathways to a given problem.
I have to admit that I can't really follow the technical discussion about blobs.
It only came to my mind that blobs are somewhat similar to electron orbitals.
Orbitals as far as I understand are clouds of probability to encounter an
electron with, if I remember correctly, increasing probability towards the
center of the cloud. This is actually quite similar to metaballs. I'm not
claiming, of course that the blobs I have been using are somehow resembling the
actual fullerene orbitals.
It was interesting to look at the read-function - I never used it... I extracted
data for small molecules always by hand from xyz-files. In the case of large
molecules I wrote my own Perl-scripts for transforming pdb-data into pov-files.
As a teaser for these larger molecules, here comes a detail from the active
center of an endoglucanase with its substrate.
Btw, I apologize for posting so rarely and that it will also take some more time
until my next video...
Thomas
Post a reply to this message
Attachments:
Download 'endoglucanasej.jpg' (150 KB)
Preview of image 'endoglucanasej.jpg'
|
 |




![]()